 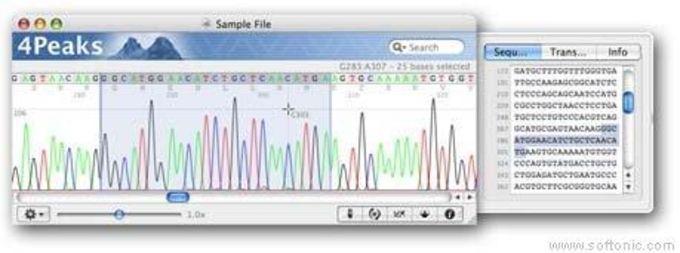
The common sequencing artifacts, which cause most standard base-calling tools to miscall the chromatogram traces, include: 1) polymerase slipage, 2) loss of resolution, 3) contamination and 4) dye blob. However, most base-calling tools developed for conventional sequencing may not be suitable for SNP detection because they usually misinterpret chromatogram traces at heterozygous base positions. The efficiency of nucleotide variation detection relies mainly on the accuracy of bioinformatic software used to base-call the chromatograms. The terms SNP and point mutation are considered synonymous for the algorithm described in this paper. Most fluorescence based sequencers produce nucleotide signals (chromatograms) that must be base-called in order to detect the SNP or point mutation. The discovery of SNPs may help to identify causative gene mutations in monogenic diseases as well as SNPs associated with predisposing genes in complex diseases. įollowing completion of the human genome project, detection and discovery of single nucleotide polymorphisms (SNPs) is at the forefront of genomic research. 
The current version of VarDetect is freely available at. VarDetect is compatible with most major operating systems such as Microsoft Windows, Linux, and Mac OSX. 
In this comparison of automatic SNP detection tools, VarDetect achieved the highest detection efficiency. These chromatograms were obtained from sequencing 16 two-pooled DNA samples a total of 32 individual DNA samples. The proposed software tool is benchmarked against four other well-known SNP discovery software tools (PolyPhred, novoSNP, Genalys and Mutation Surveyor) using fluorescence based chromatograms from 15 human genes. Accurate SNP base-calling is achieved using pre-calculated peak content ratios, and is enhanced by rules which account for common sequence reading artifacts. We present VarDetect, a stand-alone nucleotide variation exploratory tool that automatically detects nucleotide variation from fluorescence based chromatogram traces. Accurate detection of SNPs requires software that can correctly interpret chromatogram signals to nucleotides. The discovery of such variation may help to identify causative gene mutations in monogenic diseases and SNPs associated with predisposing genes in complex diseases. Single nucleotide polymorphisms (SNPs) are the most commonly studied units of genetic variation. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
